OK readers, here we go. Now that you have the basics, let’s get back to the hero of this series, my favorite enzyme, Aminoacyl tRNA Synthetase (usually referred to as aaRS, not Syzase). Remember that the task of the enzyme is to join the right tRNA with the right (cognate) amino acid. The tRNAs each carry one amino acid to the ribosome where their cargo (the amino acid) is added to the growing protein chain in the correct order. This happens because the tRNAs, using their anticodon triplets, bind in turn to the codons on the mRNA, which have been copied from the master code sheet, the DNA.
Amino acids and tRNAs cannot recognize each other because they are parts of different chemical systems, so there is no way that the correct amino acid (matching the tRNA anti codon) could ever get matched with its tRNA without help. It would be like an English-only speaker having a conversation with an Italian-only speaker. They need a translator. And the helper/translator is aaRS. That’s why I consider this enzyme to be at the heart of the entire translation process.
How does the enzyme do this? First, each enzyme has a binding site for its specific amino acid. So alanine aaRS has an alanine binding site. It also has a binding site for each of the 4 tRNAs that carry one of the 4 anticodons for alanine. Once this very large enzyme has alanine bound in one site and an alanine tRNA bound on the other site, the protein conformation (the protein’s shape) shifts in a way so that the end of the tRNA and one end of the amino acid are close. The enzyme then catalyzes a chemical reaction (requiring energy) between the two molecules, and the result is a strong chemical bond between tRNA and amino acid. The process is illustrated in the following figures.
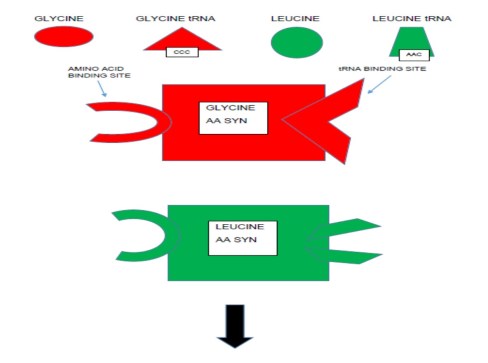
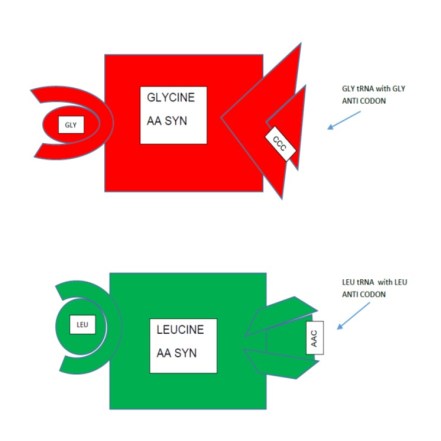
Imagine the interpreter saying “Mr. Smith, let me introduce Mr. Russo; Signor Russo, le presento il Signor Smith.” And the two men shake hands. Only in the chemical case they don’t let go; they are bound.
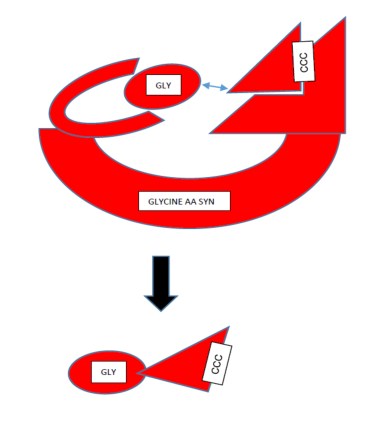
So the enzyme has done its job. It has taken its amino acid and matched it with the right tRNA so that the amino acid will be put in exactly the right place determined by the DNA sequence. Once this is done, the new bonded tRNA-amino acid complex can leave its binding sites, and as a tightly bound couple navigate its way to the ribosome, where the two molecules will part company, the tRNA to go back to finding another aaRS (or maybe the same one), and the amino acid to live the rest of its life as part of a protein.
But wait, we aren’t done. If this were the whole story, it would be amazing and exciting, but there’s more. You see, the entire process of translation really needs to be very accurate. If the wrong amino acids get attached to the wrong tRNA, the protein sequence will not be what it’s supposed to be, and the protein might not work. In fact, the entire translation process, including the work of the aaRS, is extremely accurate, with one mistake in over 10,000 trials. To reach that level of accuracy, the binding and matching I described above just won’t cut it. Many amino acids are very similar to many other amino acids (just ask a biochem grad student), and mistakes can definitely be made. A leucine might just fit into the binding site on an isoleucine aaRS, and that wouldn’t be good. And alanine is only slightly bigger than a glycine, and it would take an amazingly well engineered binding site to be able to distinguish the right from the wrong amino acid with 99.999% success.
So the enzyme has some tricks to make sure it hasn’t screwed up. Once the amino acid and tRNA are bound, the happy couple are shunted to another site on the enzyme called the editing site. This site is shaped in a way that will allow almost any amino acid and tRNA to get in, except the right ones. That particular amino acid tRNA just doesn’t fit. It’s especially good at admitting smaller amino acids while excluding the right one. So if everything is correct, the amino acid tRNA does not (cannot) bind in the editing binding site, and it breaks free from the enzyme to go on its merry way toward the ribosome. But if a mismatch or some other error had taken place, and the mistakenly bound couple does fit into the editing site, it’s cut into pieces by the action of the enzyme and never gets close to the ribosome.
So not only does my favorite enzyme recognize the proper amino acids and from 1 to 6 proper tRNAs, and not only does it provide the energy and mechanical means to link the two together, it also makes sure it got it right and deals a death blow to any erroneous products. Pretty neat, eh?
And I should mention that every cell in every living thing on this planet has the same system and has had it as far back as LUCA. In fact, we have no idea what came before and how the present universal system evolved, since it’s this system that is the biochemical key to evolution. But we can discuss this another time. For now I hope you will agree that aminoacyl tRNA Synthetase should be everyone’s favorite enzyme.

Very cool! Thank you, and I’m eagerly wondering what comes next.
Thanks Sheila. I wonder also.
Well Sy, I certainly went down the rabbit hole following this series of articles. I have an old friend who’s now an adjunct professor in Hawaii, an MIT “Cell & Molecular Biologist” who specialized in protein engineering. Probably twenty-hours into just getting a grip on the jargon… some videos helped tremendously. (“It was so large a house, that she did not like to go nearer till she had nibbled some more of the lefthand bit of mushroom, and raised herself to about two feet high…”)
I get the “interpreter” issue, and see how it’s essential to the creation of every protein (including itself)… and thus also the paradox… chicken AND egg. (“I don’t see how he can EVER finish, if he doesn’t begin.” -Alice thinks to herself.) So what *are* the odds?
For life to begin, it seems like it would be more important that the interpreter simply TELL the (or perhaps “a”) correct first part to the story, regardless of what she first reads… if she even reads anything at all. “Make ME!” said the proto-aaRS. Natural selection is otherwise a merciless editor, even as the story grows in complexity. Though, just to throw my brain into overdrive regarding the magnitude of current interpretation, it was suggested that I look up, “ATP Synthase”…!!!
Thanks for these articles, Sy. These go right along a line I’ve considered for many years. Recently having lost a bit of the wind in my sails, these are also good to ponder in the relative calm. Cheers to you.
Thanks, Kumi, you got the point, of course, so congratulations. I love the Alice analogy, and might very well steal or at least borrow it at some point. My issue is that most people, including evolutionary biologists, dont pay sufficient attention to the biochemistry that drives evolution. I think they should.